Researchers show how a tumor cell’s location and environment affect its identity
Friday, June 23, 2023
Researchers show how a tumor cell’s location and environment affect its identity
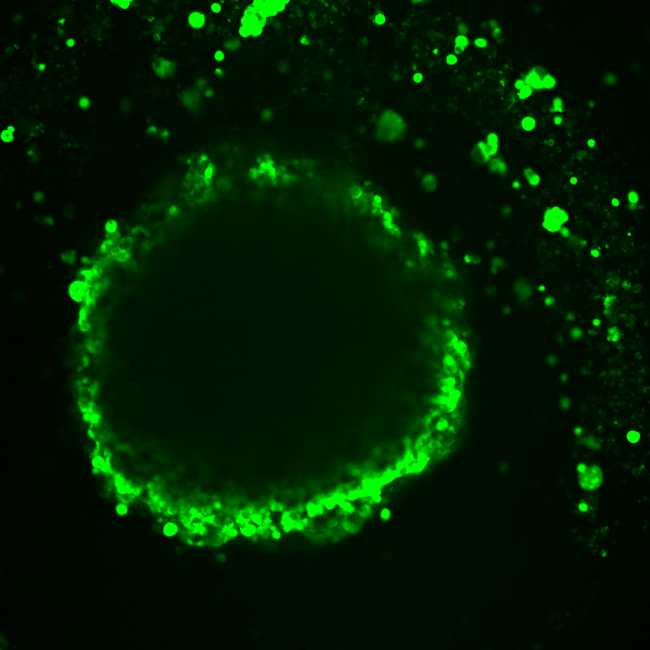
Using 3-D models of ovarian cancer tumors, scientists found differences in gene activity based on where a cell is in a tumor, demonstrating how a cell’s location and environment in a cancerous tumor can strongly influence which genes are active and the cell’s role in the cancer’s biology. More specifically, the team co-led by researchers at the National Center for Advancing Translational Sciences (NCATS), part of the National Institutes of Health, showed that gene activity in cells at or near a tumor’s surface differed from that of cells closer to the tumor center.
The approach pairs the use of a technology to reveal the genetic activity of single cells within a tumor with fluorescent dyes that spread into tumors. The work could allow researchers to study how the same diseases can vary in people and progress differently. This research could help clinicians identify treatment strategies focused on specific areas in tumors, which could lead to better therapies for cancers and other diseases. The team reported its results June 21 in Cell Systems.
“It’s commonly accepted that a cell’s location and surrounding environment influence the cell’s identity,” said Craig Thomas, Ph.D., a translational scientist at NCATS. “Two cells can be genetically identical but have different cellular identities, meaning different genes are turned on because of their location and environment. Our goal was to establish a straightforward method to study this concept in multiple settings.”
The new system, called Segmentation by Exogenous Perfusion, or SEEP, takes advantage of a dye that diffuses into cells throughout a tumor at a definable rate. Measuring how much dye gets into individual tumor cells provides information on the cell’s location, and specifically, its access to the outside environment. Using computational methods, the researchers linked this information to cells’ gene activity, allowing the scientists to connect the cells’ identities with their location.
“Understanding the relation of cells to each other and the effects of their positions in space has been a fundamental question in cancer, neurological disorders and other areas,” said co-author Tuomas Knowles, Ph.D., at the University of Cambridge.
Researchers used three types of 3-D laboratory models — spheroids, organoids and mouse models — created from human ovarian cancer cells. Spheroids are 3D clusters of cells grown in a lab dish that can mimic some traits of organs and tissues. Organoids, also grown in a dish, are more complex 3-D models that more closely mimic organ and tissue function and structure. In the mouse models, researchers implanted human ovarian cancer cells to form tumors.
“It’s critical to understand that not every cell in a tumor will be exposed to a drug in the same way,” Knowles said. “A cancer drug might kill the cells on the surface of a tumor, but the cells in the middle are different and affected differently. That’s likely contributing to why some therapies fail.”
The SEEP method revealed that tumor cells near the tumor surface were more likely to undergo cell division than cells closer to the tumor center. Cells on the surface of tumors also turn on genes to protect them from immune system responses. Not surprisingly, these gene responses are linked to how the tumor hides from the body’s immune defenses.
Researchers were surprised at the differences in gene activity between cells on or near the surface and those farther inside the ovarian cancer tumor models. The findings could help scientists better understand how tumors are structured. Such information could lead to improved treatments. One possible cancer treatment method could be to target cells likely to be affected in different areas of tumors.
“Certain tumor cell types are susceptible to certain therapies,” noted first author and Harvard University medical student David Morse, Ph.D. “Knowing where cells are located and their levels of accessibility in the tumor could help us decide how to use drugs in combination. It could help tell us how long to give a drug and when to move on to other therapies.”
The NIH research was supported by NCATS’ Division of Preclinical Innovation and the National Cancer Institute’s Center for Cancer Research. Work at Harvard University was funded by the National Science Foundation (DMR-1708729) and through the Harvard Materials Research Science and Engineering Center (DMR-2011754). The research at Cambridge University was supported by the Biotechnology and Biological Sciences Research Council, the Newman Foundation, the Wellcome Trust, and the European Research Council under the European Union’s Seventh Framework Program (FP7/2007–2013) through the European Research Council grant PhysProt (agreement no. 337969). There also was support by the NIH Oxford–Cambridge Scholars Program and the Certara Biomedical Research Scholarship.
About the National Center for Advancing Translational Sciences (NCATS): NCATS conducts and supports research on the science and operation of translation — the process by which interventions to improve health are developed and implemented — to allow more treatments to get to more patients more quickly. For more information about how NCATS helps shorten the journey from scientific observation to clinical intervention, visit https://ncats.nih.gov.
About the National Institutes of Health (NIH): NIH, the nation's medical research agency, includes 27 Institutes and Centers and is a component of the U.S. Department of Health and Human Services. NIH is the primary federal agency conducting and supporting basic, clinical, and translational medical research, and is investigating the causes, treatments, and cures for both common and rare diseases. For more information about NIH and its programs, visit www.nih.gov.
NIH…Turning Discovery Into Health®
DB Morse, et al. Positional influence on cellular transcriptional identity revealed through spatially segmented single-cell transcriptomics. Cell Systems DOI: https://doi.org/10.1016/j.cels.2023.05.003
Institute/Center
Contact
301-435-0888