Tracking SARS-CoV-2 variants in wastewater
July 26, 2022
Tracking SARS-CoV-2 variants in wastewater
At a Glance
- A system that combined advanced sample preparation with powerful computer analysis was able to identify individual SARS-CoV-2 variants in wastewater.
- Using wastewater to track the emergence of new variants of concern could be faster and less expensive than clinical testing.
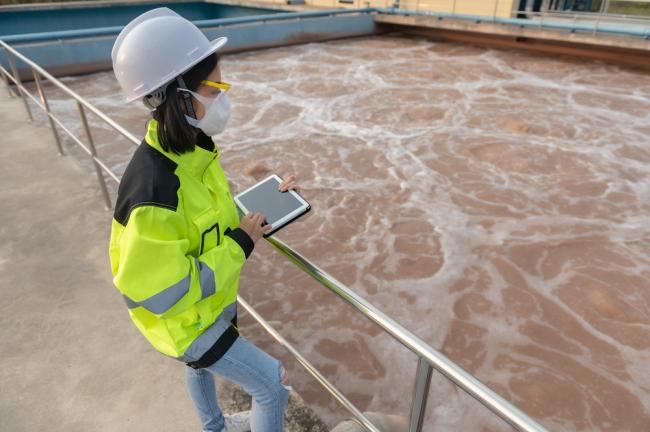

During infection with SARS-CoV-2, the virus that causes COVID-19, people shed virus down the drain every time they wash their hands or use the toilet. This can happen whether or not someone has symptoms of the disease.
Scientists have been tracking levels of SARS-CoV-2 in wastewater as a way to estimate whether infections are rising or falling in communities. Using wastewater to track viral transmission has many potential advantages over clinical testing, which is expensive and requires people to seek out testing.
To date, it’s been difficult to obtain information about specific variants of SARS-CoV-2 from wastewater. RNA, the genetic material of the virus, is easily damaged. It’s also relatively scarce in water samples. This has made it difficult to gather enough high-quality SARS-CoV-2 RNA from wastewater to identify variants rather than just detect the virus in general.
In a new study, a team of researchers from the University of California, San Diego (UCSD) and Scripps Research Institute aimed to surmount these limitations. They designed a type of nanobead that binds efficiently to viral RNA in wastewater, allowing more to be captured intact for sequencing.
The team also developed a computer analysis tool to recognize small, distinct pieces of these sequences. It could rapidly tag different viral variants in the collected samples and estimate their abundance.
The researchers tested the new monitoring system between November 2020 and September 2021. They examined wastewater gathered daily around the UCSD campus. They also examined daily samples from the primary wastewater treatment plant serving the greater San Diego County. Results from the study, which was funded in part by NIH, were published on July 7, 2022, in Nature.
Trends in the wastewater samples mirrored those from clinical testing over the study period. But the wastewater sampling system detected the Alpha, Delta, and other early variants about two weeks before they started to show up in clinical test samples. The system flagged the presence of the original Omicron variant in San Diego more than a week before it was picked up by clinical sampling in the community. This early detection occurred despite the researchers having less than 3% as many wastewater samples as clinical swabs.
The wastewater sampling continued to pick up variants in the community weeks after they no longer showed up regularly in clinical testing. It also picked up rare variants that were not found often in the clinic.
"The coronavirus will continue to spread and evolve, which makes it imperative for public health that we detect new variants early enough to mitigate consequences,” says Dr. Rob Knight from UCSD, one of the study's lead authors.
“In a lot of places, standard clinical surveillance for new variants of concern is not only slow but extremely cost-prohibitive,” adds Dr. Kristian Andersen from Scripps, who also helped lead the work. “But with this new tool, you can take one wastewater sample and basically profile the whole city.”
—by Sharon Reynolds
Related Links
- Long COVID Symptoms Linked to Inflammation
- T Cells Protect Against COVID-19 in Absence of Antibody Response
- Intranasal Proteins Could Protect Against COVID-19 Variants
- Antibodies Recognize New SARS-CoV-2 Omicron Variant After Booster
- Mandatory Masking in Schools Reduced COVID-19 Cases
- COVID-19 Immune Response Improves for Months After Vaccination
References
Wastewater sequencing reveals early cryptic SARS-CoV-2 variant transmission. Karthikeyan S, Levy JI, De Hoff P, Humphrey G, Birmingham A, Jepsen K, Farmer S, Tubb HM, Valles T, Tribelhorn CE, Tsai R, Aigner S, Sathe S, Moshiri N, Henson B, Mark AM, Hakim A, Baer NA, Barber T, Belda-Ferre P, Chacón M, Cheung W, Cresini ES, Eisner ER, Lastrella AL, Lawrence ES, Marotz CA, Ngo TT, Ostrander T, Plascencia A, Salido RA, Seaver P, Smoot EW, McDonald D, Neuhard RM, Scioscia AL, Satterlund AM, Simmons EH, Abelman DB, Brenner D, Bruner JC, Buckley A, Ellison M, Gattas J, Gonias SL, Hale M, Hawkins F, Ikeda L, Jhaveri H, Johnson T, Kellen V, Kremer B, Matthews G, McLawhon RW, Ouillet P, Park D, Pradenas A, Reed S, Riggs L, Sanders A, Sollenberger B, Song A, White B, Winbush T, Aceves CM, Anderson C, Gangavarapu K, Hufbauer E, Kurzban E, Lee J, Matteson NL, Parker E, Perkins SA, Ramesh KS, Robles-Sikisaka R, Schwab MA, Spencer E, Wohl S, Nicholson L, Mchardy IH, Dimmock DP, Hobbs CA, Bakhtar O, Harding A, Mendoza A, Bolze A, Becker D, Cirulli ET, Isaksson M, Schiabor Barrett KM, Washington NL, Malone JD, Schafer AM, Gurfield N, Stous S, Fielding-Miller R, Garfein RS, Gaines T, Anderson C, Martin NK, Schooley R, Austin B, MacCannell DR, Kingsmore SF, Lee W, Shah S, McDonald E, Yu AT, Zeller M, Fisch KM, Longhurst C, Maysent P, Pride D, Khosla PK, Laurent LC, Yeo GW, Andersen KG, Knight R. Nature. 2022 Jul 7. doi: 10.1038/s41586-022-05049-6. Online ahead of print. PMID: 35798029.
Funding
NIH’s National Institute of Allergy and Infectious Diseases (NIAID), National Center for Advancing Translational Sciences (NCATS), National Center for Complementary and Integrative Health (NCCIH), and Office of the Director (OD); U.S. Centers for Disease Control and Prevention; Conrad Prebys Foundation; National Science Foundation; San Diego County Health and Human Services Agency.