Cell atlases give detailed views of human organs
August 8, 2023
Cell atlases give detailed views of human organs
At a Glance
- Researchers created detailed cellular maps of three human organs, offering new insights into human health and disease.
- The maps are among the first of many expected from an effort that aims to uncover how cells are arranged and interact in human tissues.
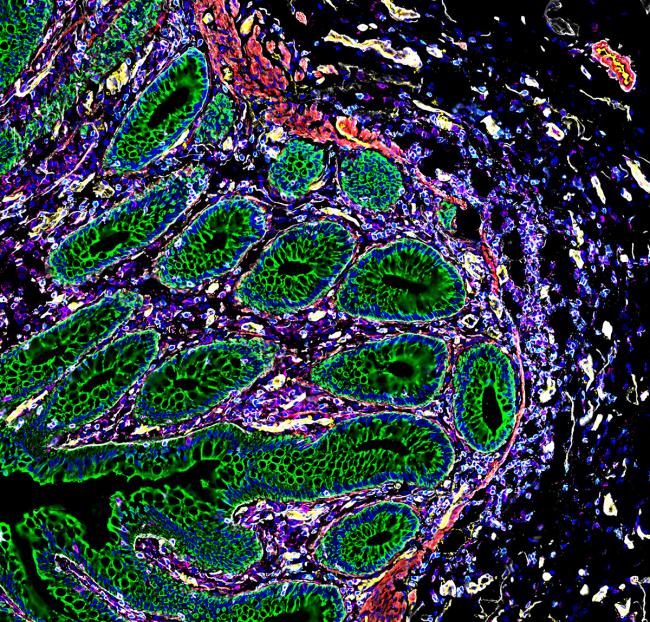
The human body comprises tens of trillions of cells. Each has specific functions. The way these cells are spatially arranged within tissues and organs can affect how they operate. How the cells interact with their surrounding environment can also play a role. Knowledge of how different types of cells function within an organ could give new insights into human health and disease. But assessing the spatial arrangement of cells within organs has been a challenge.
To better understand the inner organization of organs, NIH launched a large-scale collaborative project called the Human BioMolecular Atlas Program (HuBMAP). Since its creation five years ago, the project has supported development of research tools and techniques that can map individual cells within organs. Resulting methods and datasets are then shared with the scientific community to enable collaboration and discovery.
The project’s latest advances include the creation of cellular reference maps for three human organs: the kidney, intestine, and placenta. The maps include visual and spatial data that identify the genes and proteins that are active within cells. Results appeared in three papers published in Nature on July 19, 2023. That same date, an additional six HuBMAP-supported scientific papers were published in Cell and the Nature family of journals, describing studies related to human skin, blood vessels, and more.
The human kidney study created interactive 3D models based on analyses of 45 healthy and 48 diseased human kidneys. The highly detailed atlas shows the cellular states and locations of 51 types of kidney cells in healthy and diseased tissues. Comparative studies are expected to generate significant insights into the root causes of rare and common kidney disorders, and ultimately lead to improved treatments.
The study of the human placenta examined the maternal-fetal interface during the first half of pregnancy. The resulting atlas is based on analysis of a half-million cells and nearly 600 arteries, using samples obtained during different weeks of pregnancy. The atlas reveals how maternal arteries change and twist to embed in the placenta and supply the fetus with needed blood and nutrients. The atlas also uncovered new details about the interactions between the placenta and the mother’s immune cells.
The atlas of the human intestine details how cells are organized into distinct regions along its approximately 30-foot length. The intestine is responsible for diverse and complex activities, including digestion and maintenance of its microbiome. The researchers analyzed eight sections of the intestine from nine people. They detected new cellular subtypes and distinguished cellular communities that perform specific functions, including immune response.
“These are really some of the first highly detailed spatial maps of the human body, allowing us to see not just what cells are there, but how they’re organized,” says Dr. Michael Snyder of Stanford Medicine, a coauthor of the intestine paper and former co-chair of the HuBMAP steering committee. “It’s hard to understand disease if you don’t know what the healthy states look like. These maps will allow us to start comparing across different organs and analyzing what goes wrong during disease.”
Future HuBMAP studies are expected to create detailed cellular maps for more than a dozen additional organs. HuBMAP is part of a larger community of NIH initiatives working to create cellular maps of human tissues and organs.
—by Vicki Contie
Related Links
- Mapping the Mammalian Motor Cortex
- Building an Atlas of Brain Function in Mice
- Human BioMolecular Atlas Program
- Kidney Precision Medicine Project
- Human Tumor Atlas Network
- LungMap
References
An atlas of healthy and injured cell states and niches in the human kidney. Lake BB, Menon R, Winfree S, Hu Q, Ferreira RM, Kalhor K, Barwinska D, Otto EA, Ferkowicz M, Diep D, Plongthongkum N, Knoten A, Urata S, Mariani LH, Naik AS, Eddy S, Zhang B, Wu Y, Salamon D, Williams JC, Wang X, Balderrama KS, Hoover PJ, Murray E, Marshall JL, Noel T, Vijayan A, Hartman A, Chen F, Waikar SS, Rosas SE, Wilson FP, Palevsky PM, Kiryluk K, Sedor JR, Toto RD, Parikh CR, Kim EH, Satija R, Greka A, Macosko EZ, Kharchenko PV, Gaut JP, Hodgin JB; KPMP Consortium; Eadon MT, Dagher PC, El-Achkar TM, Zhang K, Kretzler M, Jain S. Nature. 2023 Jul;619(7970):585-594. doi: 10.1038/s41586-023-05769-3. Epub 2023 Jul 19. PMID: 37468583.
A spatially resolved timeline of the human maternal-fetal interface. Greenbaum S, Averbukh I, Soon E, Rizzuto G, Baranski A, Greenwald NF, Kagel A, Bosse M, Jaswa EG, Khair Z, Kwok S, Warshawsky S, Piyadasa H, Goldston M, Spence A, Miller G, Schwartz M, Graf W, Van Valen D, Winn VD, Hollmann T, Keren L, van de Rijn M, Angelo M. Nature. 2023 Jul;619(7970):595-605. doi: 10.1038/s41586-023-06298-9. Epub 2023 Jul 19. PMID: 37468587.
Organization of the human intestine at single-cell resolution. Hickey JW, Becker WR, Nevins SA, Horning A, Perez AE, Zhu C, Zhu B, Wei B, Chiu R, Chen DC, Cotter DL, Esplin ED, Weimer AK, Caraccio C, Venkataraaman V, Schürch CM, Black S, Brbić M, Cao K, Chen S, Zhang W, Monte E, Zhang NR, Ma Z, Leskovec J, Zhang Z, Lin S, Longacre T, Plevritis SK, Lin Y, Nolan GP, Greenleaf WJ, Snyder M. Nature. 2023 Jul;619(7970):572-584. doi: 10.1038/s41586-023-05915-x. Epub 2023 Jul 19. PMID: 37468586.
Funding
NIH’s Office of the Director, National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK), National Human Genome Research Institute (NHGRI), National Cancer Institute (NCI), National Center for Advancing Translational Sciences (NCATS), National Institute of Allergy and Infectious Diseases (NIAID), National Heart, Lung, and Blood Institute (NHLBI), National Institute of Mental Health (NIMH), and additional organizations.